The Human Genome Project is an ambitious effort to sequence every piece of human DNA. The project attracted collaborators from research institutions around the world – including the Whitehead Institute for Biomedical Research at the Massachusetts Institute of Technology (MIT) – and was finally completed in 2003.
access:
Alibaba Cloud’s June “Cloud Special” event goes live to win up to 500 yuan vouchers
Now, more than two decades later, MIT professor Jonathan Weissman and colleagues have gone beyond sequence and presented the first comprehensive functional map of genes expressed in human cells. Data from the project, published online June 9 in Cell, links each gene to its work in the cell and represents the culmination of years of collaboration on the single-cell sequencing method Perturb-seq.
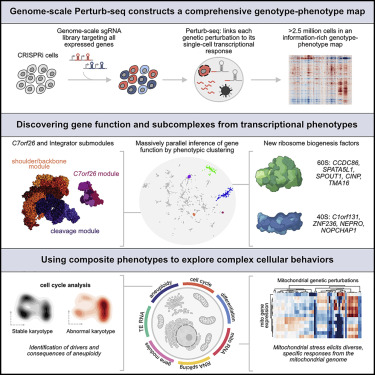
This data can be used by other scientists. “It’s a big resource, just like the human genome is a big resource, and you can go in and do discovery-based research,” Weissman said. “You don’t have to define in advance the biology you’re going to study, you have this genotype— Phenotype diagrams, you can go in and sift through the database without having to do any experiments.” He is also a member of the Whitehead Institute and an investigator at the Howard Hughes Medical Institute.
This allows researchers to delve into a variety of biological questions. They use it to explore the effects of genes of unknown function on cells, to study mitochondrial responses to stress and to screen for genes that cause chromosome loss or gain, a phenotype that has proven difficult to study in the past. “I think this dataset will enable people from other fields of biology to do all kinds of analyses that we haven’t even thought of yet, and suddenly they have this thing they can take advantage of,” said co-first author of the paper on the study , said Tom Norman, a former postdoc in Weismann’s lab.
Groundbreaking Perturb-seq method
The Perturb-seq method utilized by the project makes it possible for scientists to track the effects of turning genes on or off at unprecedented depths. This method, first published in 2016 by a team of researchers including Weissman and MIT professor Aviv Regev, can only be used for small gene sets and is expensive.
The extensive Perturb-seq map was driven by the foundational work of Joseph Replogle, an MD student in Weissman’s lab and one of the first authors of this paper. Replogle teamed up with Norman, Britt Adamson (assistant professor in Princeton’s Department of Molecular Biology), and a group at 10x Genomics to create a new version of Perturb-seq that could be scaled up. The researchers published a proof-of-concept paper in Nature Biotechnology in 2020.
The Perturb-seq method uses CRISPR-Cas9 genome editing to introduce genetic changes into cells, and then uses single-cell RNA sequencing to capture information about the RNAs that are expressed as a result of specific genetic changes. Since RNA controls all aspects of the way cells behave, this approach could help decipher the many cellular effects of genetic changes.
Weissman, Regev, and others have used this sequencing method on a smaller scale since their original proof-of-concept paper. For example, researchers are using Perturb-seq in 2021 to explore how human and viral genes interact during infection with HCMV, a common herpes virus.
In the new study, Replogle and collaborators including Reuben Saunders, a graduate student in Weissman’s lab and co-first author of the paper, scaled the approach to the entire genome. Using human blood cancer cell lines as well as non-cancer cells from the retina, he performed Perturb-seq on more than 2.5 million cells and used the data to build a comprehensive map linking genotype and phenotype.
Dig deeper into the data
After completing the screening, the researchers decided to put their new dataset to work and set out to investigate some biological questions. Tom Norman points out: “The advantage of Perturb-seq is that it allows you to get a large dataset in an unbiased way. No one knows exactly what the limits of what you can get out of such a dataset are. Now the question is, do you What is it actually used for?”
The first and most obvious application is the study of genes with unknown functions. Because the screen also reads out the phenotypes of many known genes, researchers can use this data to compare unknown and known genes and look for similar transcriptional results, which may indicate the production of these gene products as a larger complex. part work together.
Mutations in a gene called C7orf26 were particularly striking. The researchers noticed that those genes that were removed leading to a similar phenotype were part of a protein complex called Integrator, which plays a role in creating small RNAs. The Integrator complex is made up of many smaller subunits — 14 separate proteins have been suggested by previous studies — and the researchers were able to confirm that C7orf26 constitutes the 15th component of the complex.
They also found that the 15 subunits work together in smaller modules and perform specific functions in the integrator complex. “It’s not clear that these different modules are so different in function without this 1,000-foot-tall rise from the ground,” Saunders said.
And another benefit of Perturb-seq is that because the assay focuses on single cells, researchers can use the data to look at more complex phenotypes that get obscured when they are studied with data from other cells unclear. “We often take all the cells where ‘gene X’ is knocked out, average them together, and see how they change,” Weissman says, “but sometimes when you knock out a gene, you lose the same gene. Different cells behave differently, and this behavior can be ignored by the average.”
The researchers found that a subset of genes, the removal of which leads to different outcomes in different cells, is responsible for the segregation of chromosomes. Their removal causes cells to lose a chromosome or pick up an extra chromosome, a condition called aneuploidy. “You can’t predict what the transcriptional response to losing this gene will be, because it depends on the secondary effects of what chromosomes you gain or lose,” Weissman said. “We realized that we could reverse this and create this compound expression type, looking for chromosomal gains and losses. In this way, we have performed the first genome-wide screen to find the factors required for proper DNA segregation.”
“I think aneuploidy research is by far the most interesting application of this data. It captures a phenotype that you can only read out with a single cell. You can’t pursue it any other way,” Norman said.
The researchers also used their dataset to study how mitochondria respond to stress. Mitochondria, which evolved from free-living bacteria, carry 13 genes in their genome. Within nuclear DNA, about 1,000 genes are somehow related to mitochondrial function. “There has been a long focus on how nuclear and mitochondrial DNA are coordinated and regulated under different cellular conditions, especially when a cell is stressed,” Replogle said.
The researchers found that when they perturbed different mitochondria-related genes, the nuclear genome responded similarly to many different genetic changes. The mitochondrial genome, however, is much more responsive.
“Why mitochondria still have their own DNA remains an open question,” Replogle noted. “A big takeaway from our work is that one benefit of having an independent mitochondrial genome may be localized to different stressors.” or very specific genetic regulation.”
“If you have one mitochondria damaged and another damaged in a different way, those mitochondria may respond differently,” Weissman said.
In the future, the researchers hope to use Perturb-seq on different types of cells than the cancer cell lines they started with. In addition, they also hope to continue exploring their gene function map and hope that others will do the same. “This is really the culmination of years of work by the authors and other collaborators, and I’m really excited to see it continue to succeed and expand,” Norman said.